There are five different color gradients : blue, red, green, grey and yellow to red. If the edges present a weight, convert-graph is able to represent the weight of the edges by computing a color gradient proportional to edge weights and coloring the edges according to it. The option Calculate the layout of the nodes (only relevant for GML output may also be chosen, otherwise the nodes will all be in diagonal and the resulting graphic will not be very instructive. Set up to their appropriate value for the demonstration, i.e., the network will be convertedįrom tab-delimited to GML format, the source node column is 1, the target column is 2 and the weight column is 3. The form is now filled with a graph in the tab-delimited format, and the parameters have been 
In the right panel, you should now see a form entitled In the NeATmenu, select the command format conversion / layout calculation.As Cytoscape and yED are not online tools, we will only describe their utilization in the command-line section. We will explain how to display this network with NeAT, Cytoscape, yED and VisANT. 
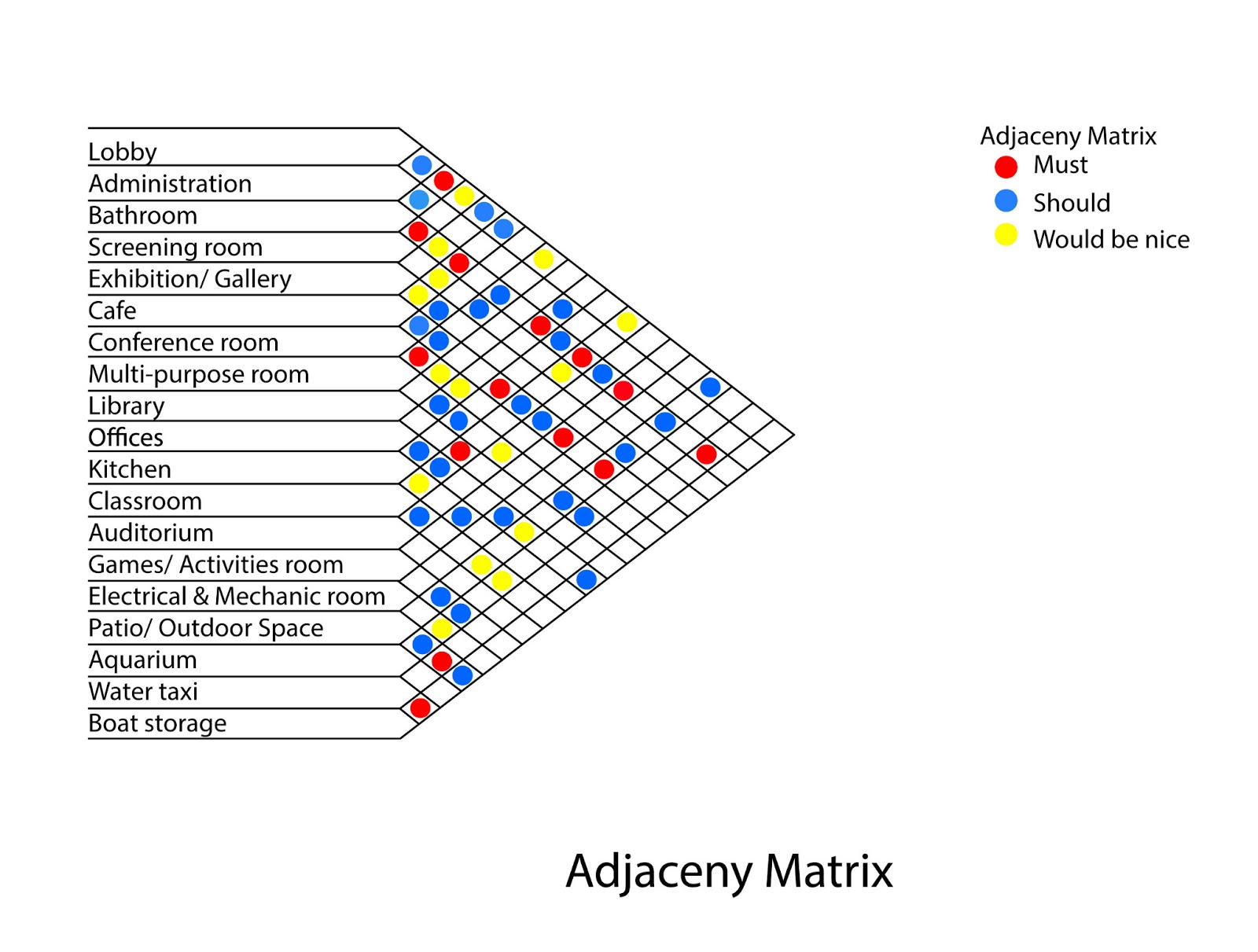
The weight expresses at which level both genes are co-expressed. An edge between two nodes means that they are co-expressed. This undirected weighted networks contains 537 nodes representing genes and 4801 edges. This network we will study consist in the top scoring edges of the yeast co-expression network included in the integrative database String. In this demonstration, we will show you how to visualize a network using some popular network visualization tools. An adjacency matrix is a x table (with the number of nodes), where a cell indicates the weight of the edge between nodes and (or 1 if the graph is unweighted).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
